APPLICATIONS
When the similar classes link is selected, a popup menu is displayed with the similar classes.
The class information along with the equivalence score is displayed for each similar class. The color
bar can contain up to three distinct colors denoting the percentage of drugs in class 1 only, class 2 only
and in both classes.
Selecting the Venn link will display the Venn diagram between the two classes,
along with the similarity table, which is a summary of clinically significant drugs
contained in each of the two classes.
Equivalence Score
Many drug classes only contain a small number of drugs, and
a small number of shared drugs between classes can
yield relatively high Jaccard values. In order to reduce
the similarity of pairs of classes with small numbers of
shared drugs, we use a modified version of the Jaccard which
we call the equivalence score.
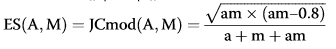
where am represents the number of drugs common to
A and M, and a + m + am the total number of unique drugs in both classes.
Example:
Class 1: Barbiturates and derivatives (N02AA)
members: Mephobarbital, metharbital, Phenobarbital, Primidone
Class 2: Phenobarbital (N0000005893) in NDFRT (has_ingredient)
members: Phenobarbital, Primidone
# common drugs: 2 (Phenobarbital, Primidone)
# unique drugs: 4 (Mephobarbital, metharbital, Phenobarbital, Primidone)
Equivalence score:
√ 2.4 /4 = 0.39
Inclusion Score
The equivalence score measures the similarity between the two classes, but does not reflect
whether one class is included in the other.
The inclusion score is a metric for finding
specific classes that are included in broader
classes. This metric combines two elements. The
first one measures the intensity of the “one-sidedness”, i.e.,
the extent to which the instances outside the intersection
are not distributed between both sides, but rather belong to
only one of the two classes. The second element measures
the coverage of the specific class by the
broader class. The inclusion score is calculated by:
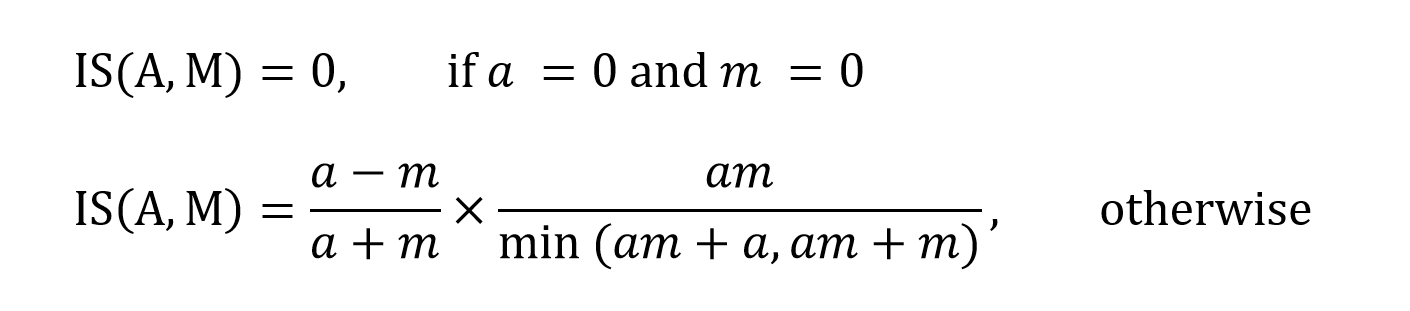
where am represents the number of drugs common to A
and M, and a and m the number of drugs specific to A
and M, respectively.
For example, if A contains 10 drugs and M contains
20 drugs and if the two classes share 9 drugs, IS(A,M) = -0.75,
providing a strong indication that A is included in M.
In summary, a negative inclusion score indicates that A is included in M, while a positive score indicates that M is
included in A. A score of 0 indicates neither set is more included. In the RxClass API, the user can return
similar drug classes for either negative inclusion scores (included_in) or positive inclusion scores (includes).